Hidden diversity found in plants’ carbon-fixing enzyme
The study suggests that plants may have more ways to tune Rubisco function than previously recognised
Rubisco is one of the quiet workhorses of life on Earth. The enzyme plays a central role in photosynthesis by catalysing the fixation of carbon dioxide, thereby driving the biochemical reactions that ultimately support plant growth and the global carbon cycle. It is also widely regarded as the most abundant protein on Earth. Yet despite its importance, scientists are still uncovering new details about how Rubisco functions in plants.
New research published in the Proceedings of the National Academy of Sciences shows that plant Rubisco is more structurally diverse than previously thought. The study reveals that different versions of Rubisco’s small subunit can coexist within the same enzyme complex in Arabidopsis thaliana, a model plant widely used in biological research.
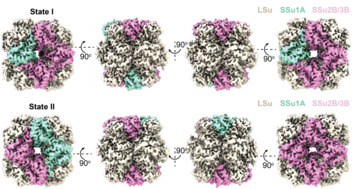
In terrestrial plants, Rubisco forms a hexadecameric complex, known as L8S8, comprising eight large and eight small subunits. The large subunit is encoded by a single chloroplast gene, whereas the small subunits are encoded by a family of nuclear genes. This enables plants to produce several related small-subunit isoforms. Until now, however, structural evidence that multiple small-subunit isoforms could coexist within individual native Rubisco complexes had been lacking.
To investigate this, the researchers purified Rubisco directly from Arabidopsis plants and analysed it using high-resolution mass spectrometry and cryo-electron microscopy. These approaches enabled the team to identify mixed-isoform Rubisco complexes and examine isoform-specific structural interactions at near-atomic resolution.
Corresponding author Dr Jani Bolla said: “Rubisco is often presented in textbooks as a single, uniform enzyme complex, but what we found is that the real enzyme in plants is more diverse than that picture suggests.”
“By combining mass spectrometry with cryo-electron microscopy, we were able to show not only that this diversity exists, but also how different isoforms contribute to the complex’s structure.”
The finding is significant because Rubisco is a major target for efforts to improve photosynthesis and crop productivity. Although the enzyme is essential for carbon fixation, it is also complex and finely balanced. Even small structural differences could affect how Rubisco performs under varying conditions, such as changes in light, temperature, or carbon dioxide availability.
The study suggests that plants may have more ways to tune Rubisco function than previously recognised. Rather than relying on a single uniform enzyme complex, plants may assemble a mixture of related Rubisco complexes with subtly different structural properties.
Dr Bolla added: “This gives us a series of snapshots of Rubisco as it occurs in plants. The next challenge is to understand what this diversity means for enzyme performance and whether plants use it to adapt photosynthesis to changing environments.”
The work provides a foundation for future research into how small-subunit isoform diversity influences Rubisco kinetics and functional plasticity. This could be particularly relevant to understanding how plants acclimate photosynthesis to changing environments and to long-term efforts to improve crop performance.




