Postdoc Poster Session 2022
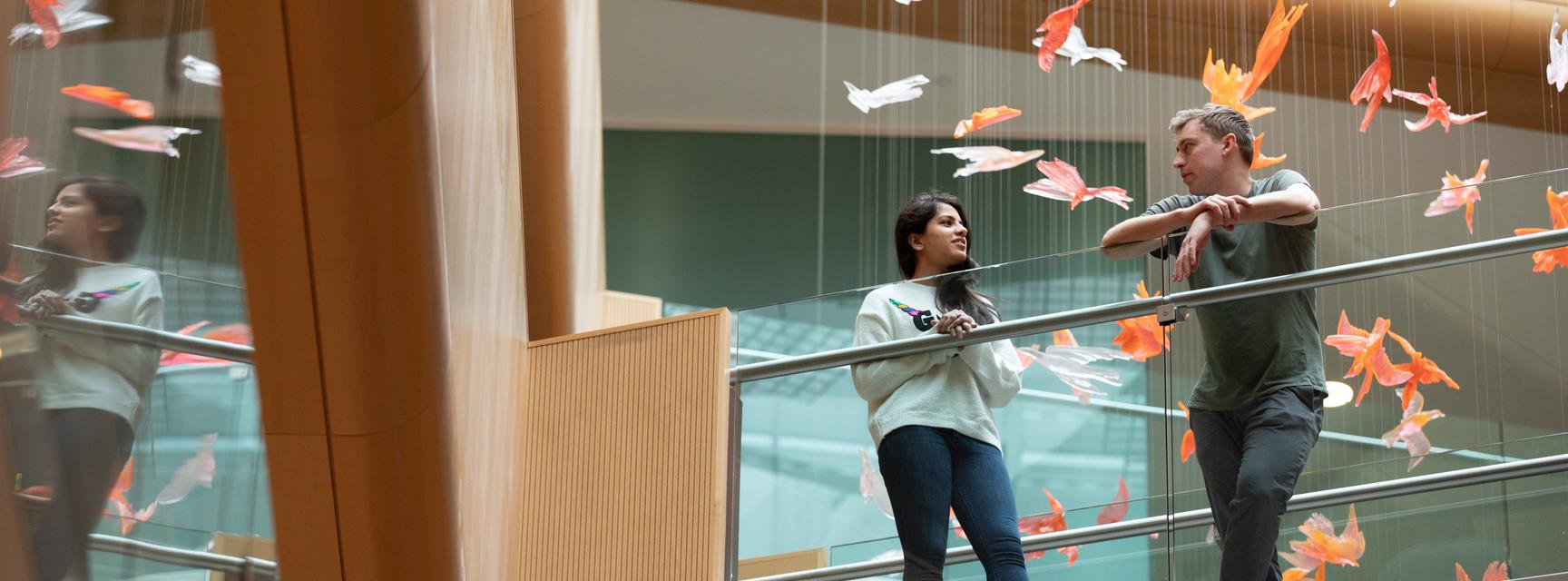
On Tuesday April 26th, the Biochemistry Postdoc Committee hosted a Poster session in the Dorothy Crowfoot Hodgkin Building. Postdoctoral researchers from the Department of Biochemistry, the Kavli Institute for Nanoscience Discovery (KIND), the Department of Physiology, Anatomy and Genetics (DPAG), the Sir William Dunn School of Pathology, and the Centre for Medicines Discovery (CMD) used the opportunity to present their research to their peers and other members of neighbouring departments. Forty-four posters were on display with additional abstracts circulated from those who were unable to present on the day. The Poster session was followed by a drinks reception allowing researchers to network and get to know others in the building to help develop a better sense of community in the building as a whole.
Read on to find out more about the research on display by researchers from the Department of Biochemistry. The full document of Poster Abstracts for participants from other departments can be provided on request from communications@bioch.ox.ac.uk
Quantitative real-time kinetic study reveals the transportation mechanism of T9SS
Xiaolong Liu
Ben Berks group
The type IX secretion system (T9SS) is a special protein transportation pathway that identified recently in bacteria of the Gram-negative Fibrobacteres-Chlorobi-Bacteroidetes superphylum and is essential for the pathogenicity of several periodontal pathogens and motility for some gliding species. The T9SS is composed of over 19 different proteins and secrets substrates that contain either a type A or type B C-terminal domain (CTD). Although the structures and interactions of some components have been revealed, it remains largely unclear how the T9SS is organized and operated. Here, we applied the split-luciferase and developed a real-time quantitative assay to study the kinetics of substrate transportation by T9SS. Our study quantitatively calculated the transportation rate of T9SS and indicated this rate variates due to the affinity of different CTDs and substrate size. This hypersensitive assay also revealed the function of some T9SS components which was not reported before. Moreover, our results indicated a competition mechanism between the transportation of type A and type B CTD substrates. Together, our study shed light on the organization and operation mechanism of T9SS and will benefit the future study on new antibiotic development.

Phenotypic manifestation of ⍺-synuclein strains derived from PD and MSA in iPSC-derived DA neurons
Benedict Tanudjojo
George Tofaris group
The progressive accumulation and subsequent aggregation of a small neuronal protein, alpha-synuclein, contributes majorly to the symptoms that characterise Parkinson's disease (PD) and related neurodegenerative disorders. To date, only partially effective symptomatic therapies exist. Therefore, there is an urgent unmet need to inform curative strategies by attempting to answer the question of how does aggregated alpha-synuclein kill neurons? To investigate the mechanisms underlying neuronal death, we generated a model using human neurons derived from patient stem cells, which recapitulated key disease features. Our study demonstrated that the extent of aggregation was dependent on the abundance of alpha-synuclein within neurons-and importantly, that the toxicity of alpha-synuclein was determined by subtle differences in the three-dimensional structure of aggregates. We screened for proteins that interacted with structurally different alpha-synuclein aggregates and discovered 56 candidates, one of which we further explored. In summary, our findings provide an insight into the complex nature of aggregation and how it ultimately leads to the demise of neurons. We define (i) alpha-synuclein levels, (ii) aggregate morphology and (iii) divergent interacting partners as the most important players in our model.
Keywords: Seeded aggregation, iPSC, Synuclein strain
Defining the cellular ZUP1 complex in genome stability
Martin AttwoodIan
Gibbs-Seymour group
The cellular response to genotoxic insults results in a complex signalling network referred to as the DNA damage response (DDR). Ubiquitin modifications on target proteins elicit several functional roles during the DDR, including the recruitment of proteincomplexes or serving as a signal for degradation. ZUP1 has recently been identified as the only member of a novel deubiquitinase enzyme (DUB) class, which has a preference for cleaving K63-linked polyubiquitin chains. It has been shown that ZUP1 can be recruited to sites of DNA damage and that ZUP1-depleted cells have DNA replication stress phenotypes. The mechanism of how ZUP1 is regulated, its substrates, and how its DUB activity shapes repair outcomes at specific DNA lesions remains unclear. Here, usingproteomic approaches, we identify a number of novel interactors of ZUP1, including the ubiquitin segregase VCP/p97 and its co-factors, various DUBs and E3 ligases. We identify the region of ZUP1 that mediates the interaction with the VCP/p97 complex and how ubiquitin-dependent signalling promotes its recruitment to ZUP1. Unexpectedly, loss of VCP/p97 function causes proteasome-dependent degradation of ZUP1, indicating a requirement for VCP/p97 in promoting ZUP1 function. Together, our work provides novel insights for understanding the cellular functions of ZUP1 in the DDR.
Keywords: DNA damage response, Deubiquitinating enzyme, Ubiquitin signalling, p97/VCP
Activation of the ZUP1 deubiquitylase by the RPA complex
Benjamin Foster
Ian Gibbs-Seymour group
Deubiquitylating enzymes (DUBs) are important regulators of ubiquitin signalling in various cellular processes, such as the DNA damage response (DDR). ZUP1 was recently discovered as the founding, and only, member of a new DUB class and was proposed to be involved in the DDR. However, our understanding of its function and the mechanistic basis for this remains in its infancy. Here, we show that the heterotrimeric RPA complex (RPA70, RPA32, RPA14), a single-stranded DNA (ssDNA) binding protein involved in DNA replication and multiple DNA repair processes, is a major interactor of ZUP1. Using both cellular and biochemical approaches, we show that the ability of ZUP1 to interact with the RPA complex is genetically encoded within ZUP1, providing a mechanism for substrate engagement. We show that the interaction between ZUP1 and RPA is direct and occurs irrespective of DNA and RPA complex phosphorylation. Moreover, we define the region in ZUP1 critical for this interaction and further show that ZUP1 interacts with both RPA70 and RPA32. Functionally, the ZUP1-RPA complex interaction dramatically activates ZUP1 K63-linked ubiquitin chain hydrolysis activity, a function which is conserved across multiple species. Together, our results suggest that the role of the ZUP1-RPA interaction is to promote ZUP1-dependent removal of ubiquitin chains from poly-ubiquitylated substrates during DNA repair processes on ssDNA.
Keywords: DUB, Ubiquitin, DNA repair, RPA
Unveiling the essential role of the exonuclease XRN1 in alphavirus replication
Vincenzo Ruscica
Ilan Davis group
The 5’-3’ exoribonuclease XRN1 is an RNA-binding protein (RBP), master regulator of RNA metabolism involved in mRNA degradation, transcription, and translational control, with proposed roles in the regulation of viral infection.The alphavirus Sindbis (SINV) efficiently infects human cells and causes a pervasive degradation of cellular RNA; at the contrary, the viral RNA (vRNA) emerges as the most stable and abundant RNA in the cell. Surprisingly, cells lacking XRN1 are refractory to SINV infection.The goal of my research is to understand whether and how XRN1 regulates SINV replication cycle.Our results reveal that re-introduction of XRN1-WT into XRN1-KO cells restores SINV viral fitness, whereas the catalytically inactive mutant fails to do so. Single molecule fluorescence in situ hybridization (smFISH) showed that XRN1 partially localize to viral replication factories, and protein-protein interaction analyses confirmed that XRN1 interacts with several viral proteins. These results highlight the importance of XRN1 for the infection of an arthropod-borne virus, and one of the most prominent viral families with biomedical relevance. Further work should be performed to decipher the mechanism by which XRN1 influences alphavirus replication and to explore the potential of the 5’-3’ degradation pathway in antiviral treatments.
Keywords: Virus, RNA, XRN1, smFISH
Peptidoglycan maturation controls outer membrane protein assembly in Gram-negative bacteria
Gideon Yaacov Mamou
Colin Kleanthous group
Linkages between the outer membrane of Gram-negative bacteria and the peptidoglycan layer are crucial to the maintenance of cellular integrity and enable survival in challenging environments. The functionality of the outer membrane relies on outer membraneproteins (OMPs), which are inserted by the β-barrel assembly machine, BAM. Growing Escherichia coli cells segregate old OMPs towards the poles by a process known as binary partitioning, the basis of which is unknown. In this study, we demonstrate that peptidoglycan underpins the spatiotemporal organization of OMPs. Mature, tetrapeptide-rich peptidoglycan binds to BAM components and suppresses OMP foldase activity. Nascent peptidoglycan, which is enriched in pentapeptides and concentrated at septa, associates with BAM poorly and has little impact on its activity, leading to preferential insertion of OMPs at division sites. The synchronization of OMP biogenesis with cell wall growth results in the binary partitioning of OMPs as cells divide. Our study revealsthat Gram-negative bacteria coordinate the assembly of two major cell envelope layers by rendering OMP biogenesis responsive to peptidoglycan maturation, a potential Achilles heel that could be exploited in future antibiotic design.
Keywords: Cell biology, Bacteria, Outer membrane, Peptidoglycan, SIM microscopy
The Spy toolbox for molecular assembly: introducing pH and temperature responsiveness
Susan Vester
Mark Howarth group
The SpyTag/SpyCatcher system has been engineered from a surface protein of the bacterium Streptococcus pyogenes as a fast and genetically-encodable tool for generating an irreversible peptide-protein interaction. Proteins in diverse biological environmentscan be empowered by SpyTag for anchoring, stretching and nanoassembly, through spontaneous formation of a covalent bond to SpyCatcher partners. We used rational engineering and phage display selection to generate a dually switchable non-covalent version of SpyCatcher, termed SpySwitch. SpySwitch allows gentle capture of SpyTagged proteins at neutral pH and release at mild acidic pH. Towards clinical testing of Spy-based vaccines, we have used SpySwitch for purification of SpyTag-and SpyTag003-fusions frombacterial or mammalian culture. We have also established thermosensitivity, so we could capture targets at 4 °C and release at 37 °C. SpySwitch-purified proteins can directly be reacted for plug-and-play nanoassembly using a panel of SpyCatcher scaffolds of diverse symmetry. Towards a vaccine with broad protection against emerging strains of SARS-CoV-2 and other coronaviruses, we have used SpySwitch to purify diverse receptor binding domains for assembly onto multimeric scaffolds.
Keywords: Protein engineering, Synthetic biology, Covid-19

Invariant surface glycoprotein 65 of Trypanosoma brucei is a complement C3 receptor important for virulence
Alex Cook
Matthew Higgins group
African Trypanosomes replicate in the blood of mammalian hosts yet are completely exposed to the adaptive and innate immune systems. Despite this, Trypanosomes can sustain long-term infections -how can this parasite survive constant host immune-surveillance? Trypanosomes evade the adaptive immune response through antigenic variation of a surface coat consisting of a dense layer of variable surface glycoprotein. However, very little is known about how they negate the innate system, including the blood circulating complement system. We have discovered that an invariant surface glycoprotein, ISG65, is a receptor for Complement Component 3 (C3). We show how ISG65 binds to the thioester domain of C3b. We also show that knockout of the ISG65 locus greatly decreases the pathogenicity of trypanosomes in a mousemodel. Deposition of C3b on pathogen surfaces is a central point in activation of the complement system and C3b has been observed on trypanosome surfaces. Our findings therefore suggest that trypanosomes have a C3 receptor distributed across their surfaces that greatly decreases their susceptibility to complement-mediated killing.
Keywords: Infectious disease, Complement system, Trypanosome, Structural biology
Direct visualisation of XistRNA in X chromosome inactivation using in situ cryogenic electron microscopy
Joy Jeongyoon Choi
Neil Brockdorff group
Xistlong non-coding RNA is the master regulator of X chromosome inactivation (XCI) process, by which one of the two X chromosomes in cells of female mammals is almost completely silenced during early development. The X chromosome is laden with many important genes for cellular differentiation as well as cancer-related genes. Accordingly, abnormal XCI has specific disease links such as muscular dystrophy and various cancers (Lalith et al., 2016).Many important studies have provided crucial insights in the XCIprocess using genetics, cell biology and genome biology approaches. However, almost all studies have been performed in denatured cells, of which resulted limited understanding in structural-functional aspect of the XCI mechanism.With recent breakthroughin in situ cryogenic electron microscopy, now is an opportune time to directly visualise many cellular processes in near-native states. Here, we employ cryo-focussed ion beam milling and correlative light and electron microscopy (cryo-FIB-CLEM) to captureX chromosome inactivation in near-native mouse embryonic stem cell. We will discuss our latest progress in this poster session.
Keywords: In situ cryoelectron cellular tomography, X chromosome inactivation
N6-methyladenosine promotes Xist RNA degradation
Guifeng Wei
Neil Brockdorff group
N6-methyladenosine (m6A) is the most abundant internal RNA modification that has been implicated in a variety of RNA processes largely through "readers" interacting with different effectors. Among thousands of m6A-decorated genes, Xist RNA that initiates Xchromosome inactivation is heavily modified at several clusters. The molecular mechanism that m6A acts on Xist RNA is however not fully understood. To dissect the immediate influence of m6A machinery on Xist and X chromosome silencing, we apply the degradation tag (dTAG) degron system to inhibit m6A pathway. We find that loss of METTL3 prolongs Xist RNA stability and in turn accelerates X chromosome silencing. Leveraging RNA-SPLIT (sequential pulse localization imaging over time) approach, we further show that prolonged Xist RNA stability has negative feedback on its RNA transcription. Depletion of m6A nuclear reader YTHDC1 or ZCCHC8, the component of the nuclear exosome targeting complex (NEXT) that interacts with YTHDC1, also lead to delayed Xist RNA degradation. Collectively, these findings suggest that m6A on Xist RNA promotes degradation.
Keywords: Xist, N6-methyladenosine, METTL3, YTHDC1, NEXT complex
Flip side of the coin: IFNg-induced trained immunity in non-immune cells
Pawel Mikulski
Lars Jansen group
Innate immunity involves an acute, mostly non-specific cellular response to infection. Unlike adaptive immunity that mounts a specific long-lasting memory of infection, the innate immune system acts transiently. However, recent insights revealed that innate immune cells such as those in the monocyte linage can maintain a "primed" state (1). Such so-called "trained immunity" allows cells to maintain a poised state following initial pathogen infection or exposure to inflammatory signals and show heighten resistance to intracellular pathogens such as viruses (2). Improperly regulated trained immunity impacts chronic inflammation in cancer, autoimmune pathologies or infectious diseases. Even though trained immunity has been described as a phenomenon at the wholeorganism level in mice and patients with a strong emphasis on immune cells, how priming works at the molecular level in is largely unknown.Central to the cellular response is a phenomenon called "long-term transcriptional memory" where pathogen signals or cytokines activate gene expression which leads to a primed-transcriptional state. Such priming results in a faster and stronger transcriptional response upon a second exposure, even following several cell division cycles in the absence of the signal, potentially underpinning trained immunity. We recently discovered a set of human innate immune target genes that are strongly primed by the cytokine Interferon gamma (IFNg) in non-immune cells (3). Following priming, in the absence of ongoing transcription, cells maintain a memory of exposure as revealed by a strong reactivation of these genes up to two weeks later, under continued proliferation. The mechanisms of priming are unknown, but, based on the evidence from other memory systems (4) and our current data, they involve changes in chromatin structure at IFN target genes. Here, I present the recent discoveries behind the identification of chromatin regulators in IFNg-induced transcriptional memory.
Keywords: Chromatin, Epigenetics, Innate immunity, Interferon, Transcriptional memory
Novel biomarkers for fatty liver disease
Bevin Gangadharan
Nicole Zitzmann group
Approximately 1 in 3 people in the UK have some degree of non-alcoholic fatty liver disease (NAFLD) and most affected individuals are unaware they have the disease. Currently, the reference standard for assessing NAFLD is the liver biopsy, aninvasive, painful procedure which can be unreliable and costly. Consequently, there is a need for improved, minimally-invasive methods of determining stages of NAFLD in patients. Mass spectrometry (MS) and two dimensional gel electrophoresis (2DE) were used to separate proteins in serum from patients with NAFLD. Several potential biomarkers were identified and both Western blotting and targeted MS were used to check these biomarkers. An antibody-free assay was developed for one of our most promising NAFLD biomarkers (apolipoprotein F) using parallel reaction monitoring. Our biomarkers look promising when compared to other NAFLD biomarkers currently used in the clinic. The novel biomarkers would allow clinicians to identify NAFLD patients so they could be advised about lifestyle changes, such as diet and exercise, which could help to reverse the disease and thus save money for healthcare providers. The technology could be used to diagnose, prognose or monitor the progression of NAFLD in an individual thereby reducing the need for a liver biopsy.
Keywords: Proteomics, Mass spectrometry, LC-MS, Biomarkers, Fatty liver disease
PRC1 exerts transcription repression through regulation of long-lived promoter inactive states
Aleksander Szczurek
Rob Klose group
Polycomb Repressive Complex 1 (PRC1) has a profound effect on transcription control of thousands of genes in mESCs rendering them largely non-expressed. How this effect is achieved by PRC1 and associated H2AK119ub modification is presently not fully understood and interestingly has not yet been studied in living cells. In order to explore the role of PRC1 in transcription regulation we studied dynamic transcription of PRC1-target genes in living cells at a single allele level with single transcript sensitivity. We characterized the effect of PRC1 on transcription through rapid depletion of Ring1B and H2AK119ub to discover that it regulates long-lived transcription competence rather than directly shapes transcription bursting. Further, using single particle tracking (SPT) we studied dynamic chromatin binding of endogenously HaloTag'ged proteins responsible for regulating crucial transcription steps including transcription initiation and RNA polymerase II pause release. SPT combined with rapid PRC1 depletion shed new light on the role of PRC1 in constraining specific transcription regulatory steps, therefore, furthering our knowledgeon the mechanism of achieving gene repression by Polycomb systems.
Keywords: Live-cell transcription imaging, Polycomb, PRC1, Repression, Mouse embryonic stem cells

Viral vectored delivery of monoclonal antibody genes against blood-stage malaria
Martino Bardelli
Simon Draper group
A significant challenge in the development of a malaria vaccine is the lack of antigen formulations that induce the high concentrations of functional antibodies required to achieve protection. Using potent monoclonal antibodies (mAbs) as prophylactics, combined with anti-malarial drugs, could bypass the need for a vaccine to aid in current malaria elimination programmes, but the high cost of this technology makes it incompatible within a developing world context. Vectored immunoprophylaxis (VIP) could provide a cost-effective strategy to implement this vision. This approach uses viral vectors such as adeno-associated virus (AAV) to deliver mAbs that are expressed in situ into the plasma from virally-transduced cells following intramuscular immunisation.
Keywords: Monoclonal antibodies, Malaria, VIP, AAV
Adaptation delay causes a burst of mutation in bacteria responding to oxidative stress
Valentine Lagage
Stephan Uphoff group
The relative timing of phenotypic and genetic changes is critical for the adaptation of bacteria to stress conditions. Rapid phenotypic responses allow cells to survive transient stresses whereas population survival during severe stress relies on evolutionary rescue via the emergence of adaptive mutations. These mutations may be present as standing genetic variation or generated de-novo during stress. Here, we explored how mutation rates change in real-time when bacteria become stressed and how these dynamics relate to the underlying molecular mechanisms of adaptation. We employed a range of single-molecule and single-cell microscopy assays to image adaptation of Escherichia coli during continuous hydrogen peroxide treatment. We observed that sudden exposure to H2O2 causes a transient burst in mutagenesis. We show that oxidative stress signalling by the transcription factor OxyR is fast. However, the short time delay before H2O2 scavenging enzymes are induced leaves cells vulnerable to the formation of DNA-damaging hydroxyl radicals via the Fenton reaction. Similar observations for cells experiencing alkylation stress show that mutagenesis bursts are a general phenomenon associated with adaptation delays and may have a beneficial role in accelerating mutation supply to facilitate evolutionary rescue.
Keywords: Bacteria, DNA repair, Oxidative stress, Mutagenesis, Microscopy
Monoclonal antibodies against Plasmodium falciparum RH5-Interacting Protein (RIPR) inform blood-stage malaria vaccine design
Barney Williams
Simon Draper group
The development of blood-stage malaria vaccines has been hampered by a lack of conserved antigens capable of inducing cross-strain neutralising antibodies. The most promising blood-stage candidate to date is RH5, currently being evaluated in clinical trials. RH5 forms the RCR complex with Plasmodium falciparum RH5-interacting protein (RIPR) and cysteine-rich protective antigen (CyRPA). RIPR is a ~120kDa protein comprising 10 Epidermal Growth Factor (EGF)-like domains. We generated anti-RIPR polyclonal antibody (pAb) by immunising rabbits with full-length RIPR protein. This pAb effectively inhibited cultured parasites in a standardised Growth Inhibition Activity (GIA) assay. Recombinant RIPR protein fragments were used to reverse GIA of the anti-RIPR pAb. Using GIA reversal we determined that the neutralising epitopes are located on a 184 amino acid region of RIPR corresponding to RIPR EGF(5-8). The importance of EGF(5-8) was confirmed by the generation of GIA positive monoclonal antibodies (mAbs) only against this region. All neutralising mAbs were able to bind, but not disrupt the RCR complex. In contrast, non-inhibitory mAbs could prevent in vitro formation of the RCR complex indicating that disrupting complex formation is not the mechanism of parasite invasion inhibition. These data were used to design next-generation vaccines containing the RIPR antigen.
Keywords: Malaria, Vaccine, Antibody

Automating Single-Molecule Tracking Experiments with Python
Amy Moores
Stephan Uphoff group
Single-molecule tracking exploits the principles of single-molecule localisation microscopy to detect individual fluorescently-labelled proteins as they diffuse through live cells. These experiments are routinely carried out in the Uphoff group, particularly for investigating the effects of DNA-damaging agents and antibiotic treatments on the mobility of specific proteins within E.coli cells.Besides the clear advantages of increasing consistency in imaging and improving user-friendliness, there are many benefits to automating the acquisition of single-molecule tracking datasets. Notably, automation unlocks the capability of time-continuous measurements ofdiffusion over extended periods of time; such experiments are currently not possible because a freshly-labelled sample is required approximately 2-3 hours into the imaging session, drawing the researcher away from the microscope back to the bench for sample preparation. By automating microscopy, the conflict for researcher's time is removed, eliminating any time-gaps in diffusion measurements.Here I demonstrate full microscope control using Python. Single-molecule tracking experiments have been scripted, enabling datasets to be acquired from pre-selected regions at the press of a single button. Datasets acquired automatically have led to faithful replications of diffusion histograms obtained previously through manual acquisition, indicating that the automated program performs as accurately and reliably as our traditional manual single-molecule tracking approach.
Keywords: Single-Molecule, Microscopy, Python, Automation, Fluorescence
Integrative modelling enables mechanistic insight into kinase regulation within the E. coli chemotaxis core signalling complex
Keith Cassidy
Mark Sansom group
Chemotactic bacteria possess highly-organized protein arrays that sense and process environmental chemical signals to ultimately regulate cell swimming pattern. Despite the ubiquity and physiological importance of such arrays, many important questions surrounding the molecular details of signal transduction remain unanswered, owing especially to the lack of a high-resolution characterization of array structure. Here, I present recent integrative modeling work, combining AlphaFold2-predicted structures with new cryo-electron tomography data from lysed E. coli cells, resulting in an all-atom model of the array’s functional building block, the core-signalling unit (CSU). Taken together, these data provide new insight into the structure and organization of the comprising transmembrane chemoreceptors and histidine kinase, enabling important structural updates to the current mechanistic model of receptor-mediated kinase regulation within the CSU.
Keywords: Bacterial chemotaxis, Signal transduction, Integrative modelling, AlphaFold2, Cryo-electron tomography
Distinct roles for CKM-Mediator in controlling Polycomb-dependent chromosomal interactions and priming genes for induction
Emilia Dimitrova
Rob Klose group
Precise control of gene expression underpins normal development. This relies on mechanisms that enable communication between gene promoters and other regulatory elements. In embryonic stem cells (ESCs), the cyclin-dependent kinase module Mediator complex (CKM-Mediator) has been reported to physically link gene regulatory elements to enable gene expression and also prime genes for induction during differentiation. Here we discover that CKM-Mediator contributes little to 3D genome organisation in ESCs, but has a specific and essential role in controlling interactions between inactive gene regulatory elements bound by Polycomb repressive complexes (PRCs). These interactions are established by the canonical PRC1 (cPRC1) complex but rely on CKM-Mediator, which facilitates binding of cPRC1 to its target sites. Importantly, through separation of function experiments, we reveal that this collaboration between CKM-Mediator and cPRC1 in creating long-range interactions does not function to prime genes for induction during differentiation. Instead, we discover that priming relies on an interaction-independent mechanism whereby the CKM supports core Mediator engagement with genepromoters during differentiation to enable gene activation.
Keywords: Genomeorganisation, Mediator, Polycomb, Gene expression




